Research |
OVERALL GOAL: The elucidation of novel tumor markers for the diagnosis, prognosis, and therapy of cancer. Current projects: (i) The Smad signaling connection to cancer metastasis. (ii) The role of p53 in genome stability. (iii) Genetic susceptibility to lung cancer. Our overall research strategy is to use cancer genomics, employing primarily breast and lung cancers as model systems, to invent new tools for the diagnosis, prognosis, management and therapy of cancer.
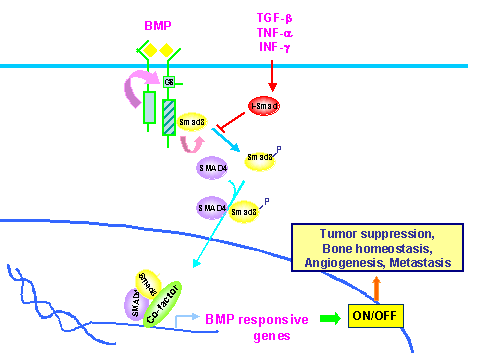
Figure 1. A model for the Smad8 connection to cancer. BMP signaling is implicated in tumor suppression, bone homeostatsis, angiogenesis and metastasis. There are numerous other signaling pathways such as the Ras-MEK pathway that could also modulate the end effects by establishing cross talk among different pathway members. Disabled Smad signaling in cancer has become increasingly recognized as an important step that affects processes such as loss of growth inhibition, promotion of angiogenesis and metastasis and the epithelial mesenchymal transition. Although frequent alterations in SMAD4 have been primarily reported in pancreatic and gastrointestinal cancers, the nature of defects involving the Smad signaling pathyways has been elusive in other cancers potentially due to alternate mechanisms and/or targets which become inactivated in the signaling pathway. Recently, we have developed a novel technique known as Targeted Expressed Gene Display (TEGD). TEGD allows for the identification of related members of a large family of genes, as well as their variants and also enables the determination of their patterns of expression in tissues and tumors. Furthermore, we extended the practical application of this technique in cancer diagnosis by analyzing the SMAD genes in cancer. We were able to carryout simultaneous evaluation of the existence as well as the levels of expression of the various Smad family members and their variants in cancers. These analyses were instrumental in providing the first clues that the loss of SMAD8 expression is assoiciated with mulitple types of cancers including 31% of both breast and colon cancers. We extended these studies to demonstrate that epigenetic silencing via DNA hypermethylation of the CpG islands in the promoter region of the gene is responsible for the loss of SMAD8 expression. The loss of Smad8 expression via epigenetic silencing in a third of breast and colon cancers makes it a significant novel tumor marker with implications for detection, prognosis and therapy of these major cancers. We hypothesize that Smad signaling downstream of the BMPs involving Smad8 could be an important pathway in metastasis/bone metastasis in cancer. As a follow up of this study, we are currently investigating the temporal relationship between epigenetic inactivation of the SMAD8 gene and the stage(s) of cancer and plan to determine the identities and roles of Smad8 signaling mediator, regulatory and effector genes in the genesis and metastasis/bone metastasis of cancer.
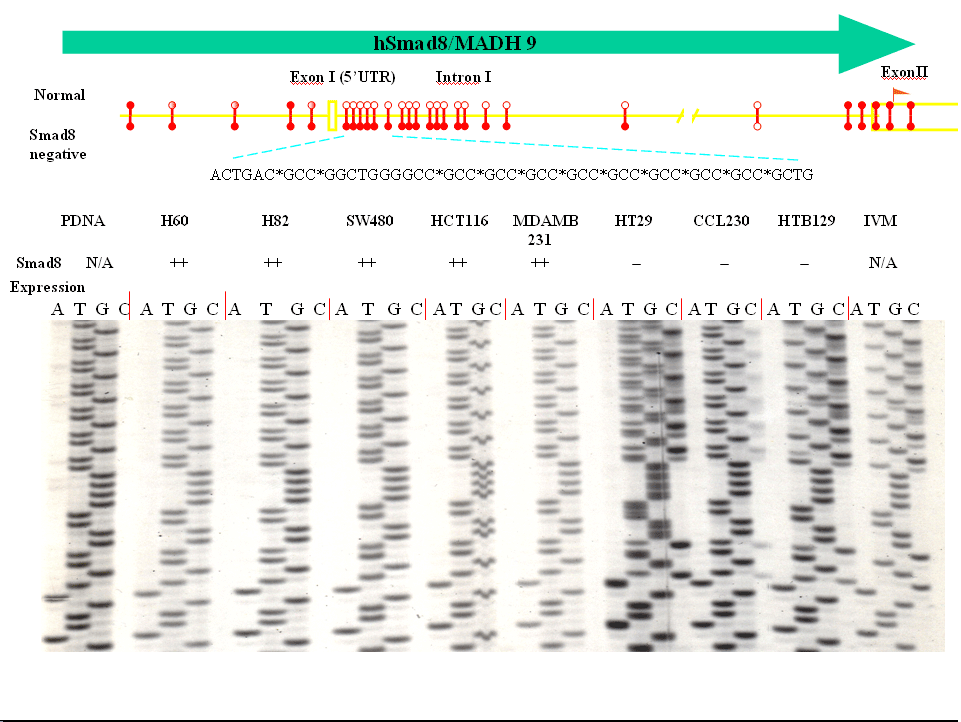
Figure 2. hSmad8 expression and promoter region GC methylation pattern. (Cheng K. et. al, Cancer Res. 64 : 1639-1646. 2004.) Our second major interest is to further the understanding of alternate modes of inactivation of p53. Because of the central nature of p53 function at the crossroads of cell death and survival, we believe that an effective therapeutic strategy based on p53 requires the understanding of the functionality of this gene product in tumors without a mutation in p53. We hypothesize that loss of p53 mediated signaling in tumors without mutations in p53 might result from a defect in an upstream event that is essential for the activation of p53 to a functional form rather than a mutation in p53. We have isolated candidate genes that modulate p53 function using a yeast double selection system, and their characterization is in progress. One of the clones that we have identified in the screen as an upstream modulator of p53 is hBUB1. BUB1 was originally identified as a spindle assembly checkpoint gene in yeast and harbors mutations in a subset of human tumors. Our indentification of hBUB1 as a modulator of p53 provided a molecular basis for the ability of p53 to maintain genomic stability. Our experimental data strongly suggest that the hBUB1 kinase is a regulator of p53 activity during the mitotic checkpoint, and inactivation of either p53 or hBUB1 in tumors could be a major mechanism for aneuploidy. In the long-term, our novel findings that link the hBUB1 and p53 genes in the maintenance of genomic stability could provide a basis for the design of novel diagnostic and therapeutic strategies for all cancers. We are also planning to study the molecular basis of p53 modulation by the other genes that we have identified. In addition to following the leads derived from existing literature on lung cancer genetics, we are also interested in understanding the molecular basis of the genesis of sporadic lung cancer caused by tobacco smoke derived carcinogens. We have formulated several experimental strategies to define the genetic and epigenetic determinants for individual susceptibility to epithelial cell DNA damage in smokers. We believe that the findings from these studies will enable the localization of hot spots for genetic alterations on the genome and hence accelerate the identification of genes targeted for inactivation during tobacco smoke derived carcinogen induced lung cancer. In the long-term, these target genes may serve as nodal points for therapeutic intervention, diagnosis, prognosis and management of the disease.
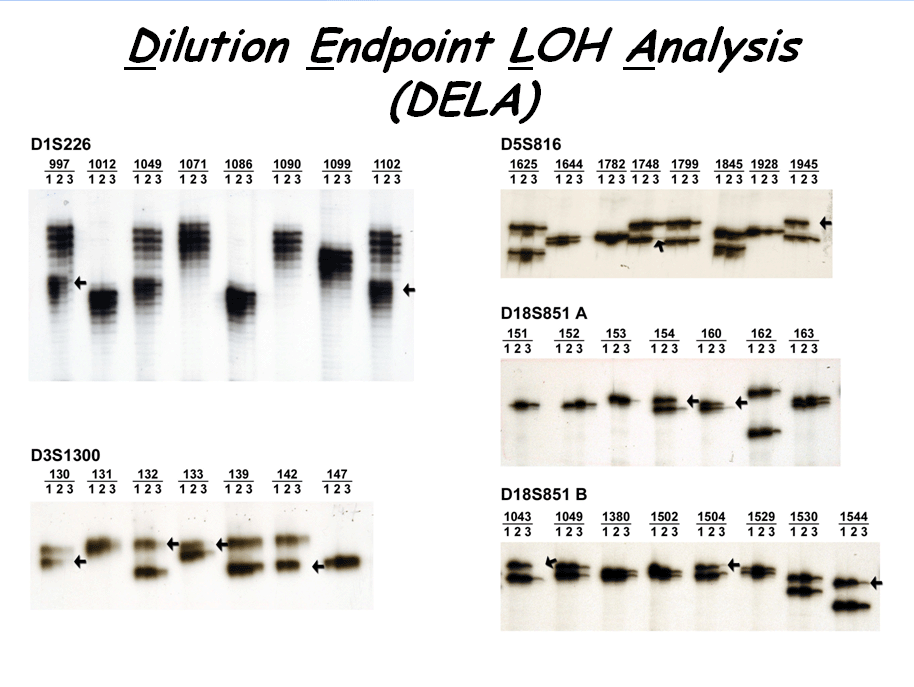
Figure 3. Lung cancer LOH detection by DELA technique. ( Pan H. et al,
Cancer Res. 65 : 1664-1669. 2005) |
|